21. TC-RMW Cross-section Plot
21.1. Description
Generate cross-section plots for TC-RMW data based on height levels using output from the METcalcpy vertical interpolation module, vertical_interpolation.py:
https://metcalcpy.readthedocs.io/en/develop/Users_Guide/vertical_interpolation.html
21.2. Example
21.2.1. Sample Data
Create the sample data by performing the conversion from pressure levels to height levels by following the instructions:
https://metcalcpy.readthedocs.io/en/develop/Users_Guide/vertical_interpolation.html
The output directory you specified in the height_from_pressure_tcrmw.sh shell script (in the METcalcpy source code) is where your tc_rmw_example_vertical_interp.nc file will be located. The tc_rmw_example_vertical_interp.nc is the input file to the cross section plot, plot_cross_section.py.
21.2.2. Configuration Files
An example configuration file (YAML, with .yaml extension) is available as a starting point for customizing the cross-plot of the height level data:
$METPLOTPY_BASE/metplotpy/contributed/tc_rmw/plot_cross_section.yaml where $METPLOTPY_BASE is the location where you saved the METplotpy source code.
# Sample YAML configuration file for generating cross-section plots
# based on height (from vertical interpolation via METcalcpy module
# vertical_interpolation).
vertical_coord_name:
'lev'
field:
'TMP'
contour_level_start: 0
contour_level_end: 500
contour_level_stepsize: 5
# time slice of interest
index_time_slice: 0
#
# contour plot appearance customizations
#
line_width: 2
contour_line_colors: 'darkblue'
contour_label_color: 'orange'
contour_label_fmt: '%1.0f'
# axis labels
# x-axis is in units of RMW
x_label: 'units of RMW'
y_label: 'Height (meters)'
# x-ticks
x_tick_start: 1
x_tick_end: 20
# y-ticks
y_tick_start: 0
y_tick_end: 3000
y_tick_stepsize: 100
# y-axis limits
y_lim_start: 0
y_lim_end: 6000
# y_scale
#y_scale: 'symlog'
y_scale: 'linear'
# plot title
plot_title: 'Temperature Cross-section'
# plot name (only provide the filename without the extension,
# png and pdf files will be generated)
plot_filename: 'example'
# plot resolution in dpi (dot per square inch)
plot_res: 300
# plot size in inches width, height
plot_size_width: 8
plot_size_height: 4.5
In this example, the plot_cross_section.yaml configuration file is set up to generate the cross-section plot for the temperature field:
- field:
‘TMP’
You can modify the plot_filename to give your output plot a different name. Two different file types are generated, a .png file and a .pdf file. In the plot_cross_section.yaml configuration file, the plot that will be created will be named example.png and example.pdf.
There are configuration settings to set the labels to the x-axis and y-axis, the plot size, plot resolution, contour colors, etc.
To generate a cross-section plot for a different field, replace the ‘TMP’ with the any other available field name from the input file (tc_rmw_example_vertical_interp.nc).
21.3. Run from the Command Line
You can generate a cross-section plot using the test_plot_cross_section.sh Bourne shell script:
# ============================*
# ** Copyright UCAR (c) 2020
# ** University Corporation for Atmospheric Research (UCAR)
# ** National Center for Atmospheric Research (NCAR)
# ** Research Applications Lab (RAL)
# ** P.O.Box 3000, Boulder, Colorado, 80307-3000, USA
# ============================*
export datadir=/path/to/vertically-interpolated-input-data
export plotdir=/path/to/output/plots
export filename=tc_rmw_example_vertical_interp.nc
export configfile=/path/to/configuration-file
python plot_cross_section.py \
--datadir=$datadir \
--plotdir=$plotdir \
--filename=$filename \
--config=$configfile
Open the test_plot_cross_section.sh script in an editor of your choice.
Modify the datadir, plotdir, filename, and configfile values in the test_plot_cross_section.sh script to point to the appropriate locations:
datadir is the location of the input netCDF file (the directory where you saved the output from the METcalcpy vertical interpolation)
plotdir is the location of the output plot
filename is the name of the input netCDF file (tc_rmw_example_vertical_interp.nc that was created from running the METcalcpy vertical interpolation)
configfile is the YAML config file that is used to set plot customizations (line colors, labels, etc.)
Save and close the test_plot_cross_section.sh script.
To generate the plot, perform the following:
- Make sure you have the following Python packages installed:
Python 3.7 or above
METcalcpy (use the same version number as this METplotpy)
matplotlib 3.4.3
metpy 1.1.0
netcdf4 1.5.7 or above
numpy
pandas
pint 0.17
xarray
yaml
Specific version numbers are specified when necessary. If versions are not specified, use a compatible version number for your operating system and existing packages.
If you are running in a conda environment, verify that you are running the conda environment that has the above Python packages installed.
cd to the $METPLOTPY_BASE/metplotpy/contributed/tc_rmw directory
cd $METPLOTPY_BASE/metplotpy/contributed/tc_rmw
replacing the $METPLOTPY_BASE with the directory where you saved the METplotpy source code.
run the test_plot_cross_section.sh script:
sh test_plot_cross_section.sh
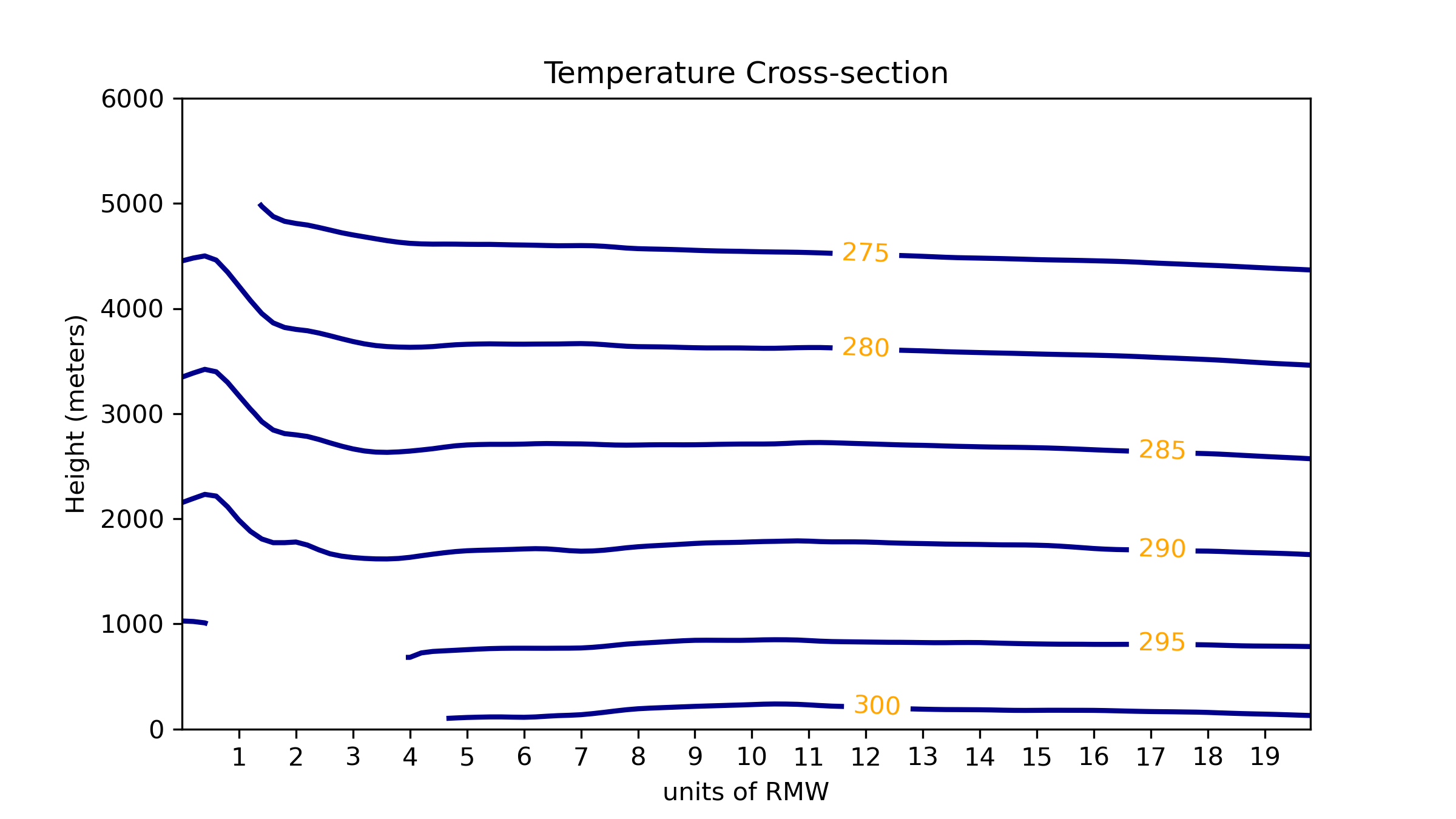
Two files will be created, example.png and example.pdf (if you used the plot_filename value of ‘example’ in the plot_cross_section.yaml configuration file). They will be located in the output directory you specified in the test_plot_cross_section.sh Bourne shell script. The plot will look like the following: